Why You Need to Prepare Reads
We previously determined that 10,000 reads gives ideal results. There is no appreciable improvement with read counts above this. After we adapter trim and quality trim the raw input reads, we select a random subsample of 10,000 reads from the remaining reads.
Although only 10,000 reads are necessary, most sequencing runs will produce many times this. To avoid overloading our servers and minimize the time for uploading, we are asking that you do an initial trimming. Taking the first 10,000 raw sequences is the simplest option, but not the most robust as there are edge effects and many of those reads could be poor quality.
Our solution is to use the first 1 million reads, adapter and quality trim, then generate the random subsample. Running simulations, there is a negligible difference in the results when randomly sampling from all sequencing reads compared to sampling from the first 1 million.
See below for instructions on how to prepare the raw reads based on your computer's operating system.
Mac OSX
- Open up Terminal. Terminal is an application located under "Utilities".

- Type cd then space.
- Now in Finder, go to the location of the FASTQ file.

- Drag the folder into Terminal. Press enter.

- Type ls then enter to verify the FASTQ is there.
- Now we want to trim this file to the first 1 million reads.
Each read is 4 lines, so we need the first 4 million lines.
Type head -n 4000000 my_file.fastq > my_file_1000000.fastq
where my_file.fastq is the name of your FASTQ file.
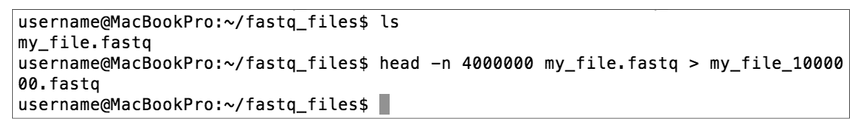
- This new FASTQ file should be less than 500 MB. If not, replace 4000000 with a smaller number until the new file is small enough.
Windows
Information coming soon...
Phred Scores
Currently, we can only handle Phred-33 scores. Sequencing done recently (after 2009) use Phred-33, so there is no need to convert the scores. For older data that uses Phred-64 (Illumina 1.5), you must convert to Phred-33 before uploading. Failure to do so will cause in poor results.
